
Neurodesk: a reproducible neuroimaging platform
18 April 2024One of our main missions at INCF is to champion the core tenets of FAIR neuroscience; we believe the fastest and most effective way for neuroscience to advance is to make disseminated brain data more Findable, Accessible, Interoperable, and Reproducible. That’s why we are always excited by new initiatives and efforts which are created in the spirit of improving FAIRness, reducing infrastructural redundancies, and mitigating analytical overlap. One project that has particularly caught our attention in such efforts is the web-based data analysis environment Neurodesk (neurodesk.org), which emphasizes portability and reproducibility in neuroimaging analysis1.
Within the neuroimaging community over the last decade, the advent of open science efforts have made possible a proliferation of both small and large-scale contributions to open source code, domain-specific data analysis packages, and software suites. Successful implementation of these tools can skirt around some of the biggest challenges in neuroimaging analyses facing small labs or research groups, most notably the financial burden incurred by subscribing to or purchasing proprietary software platforms, as well as the computational cost of conducting analyses on large-scale, multidimensional brain datasets.
However, despite the pressures alleviated by using decentralized analytical or imaging tools, new obstacles can arise along the neuroimaging analysis pipeline precisely because of the often uncoordinated and distributed nature of this software landscape. Given the numerous and often small scale of the software managers (e.g., individual researchers or labs), it is often difficult to keep track of project deprecation, cross-platform dependencies, etc. This problem is compounded by the finding that when analyzing neuroimaging data across different computing environments, systematically divergent results may be obtained even when researchers have access to the original data, code pipeline, and software versions2-4. This deals a major blow to reproducibility, one of the core principles of any scientific discipline.
Neurodesk aims to address these issues by offering a web-based platform which includes a virtual desktop (Neurodesktop), command-line interface (Neurocommand), compatibility with computational notebooks (Google Colab and Jupyter Notebooks), and leveraging software containerization (stored in their own Neurocontainers repository)5. Since containers isolate dependencies, Neurodesk offers a way for researchers to transition between software versions without the typical hassle and headache. Similarly, by making standardized analysis environments accessible with the necessary software already installed, the Neurodesk developers seek to minimize the time normally wasted on technical troubleshooting, allowing neuroscientists to put their attention and energy into the already complex tasks of their research.
With their community-oriented and open-source approach, Neurodesk is one of the newest installments in a neuroscience infrastructure ecosystem which aims to lessen the technical, legal, and financial burdens often encountered by neuroscientists, by providing an accessible and portable data analysis environment.
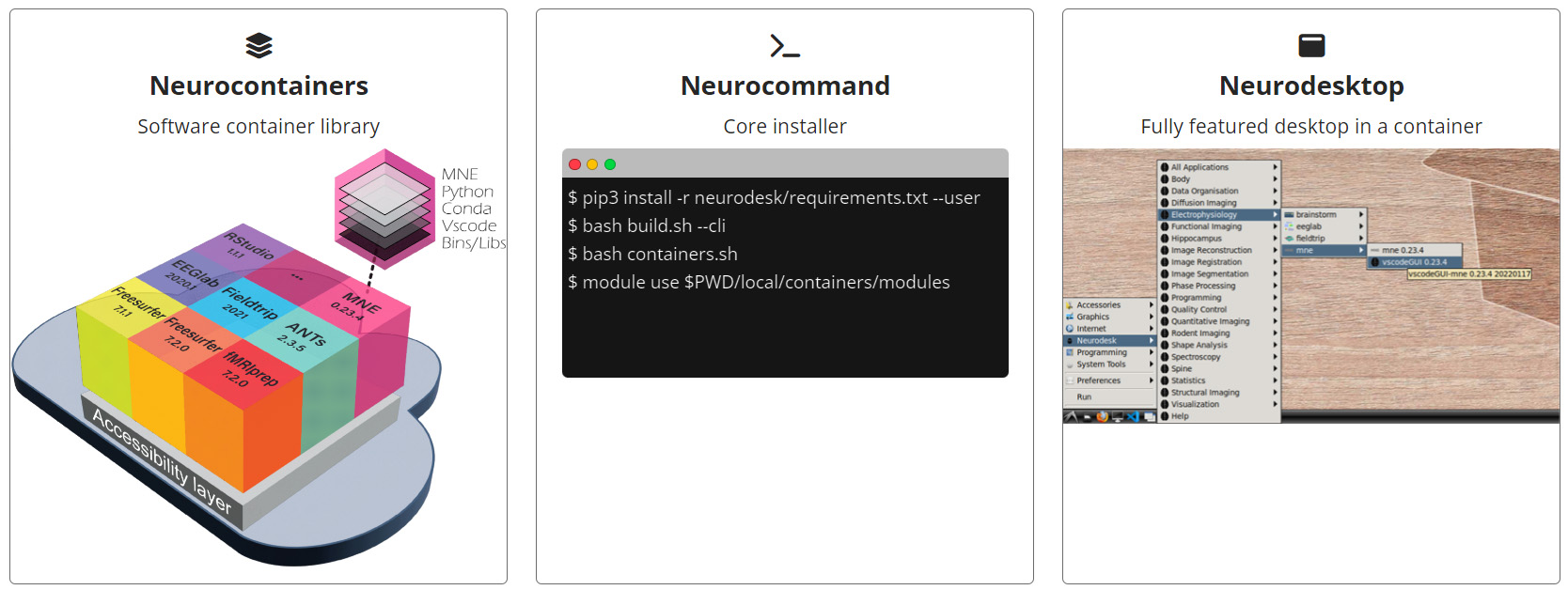
Image Credit: www.neurodesk.org
References
- Uddin, L.Q. Accessible computing platforms democratize neuroimaging data analysis. Nat Methods (2024).
- Glatard, T. et al. Reproducibility of neuroimaging analyses across operating systems. Front. Neuroinform. 9, 12 (2015).
- Gronenschild, E. H. et al. The effects of FreeSurfer version, workstation type, and Macintosh operating system version on anatomical volume and cortical thickness measurements. PLoS ONE 7, e38234 (2012).
- 21. Krefting, D. et al. Reliability of quantitative neuroimage analysis using freesurfer in distributed environments. In MICCAI Workshop on High-Performance and Distributed Computing for Medical Imaging (2011).
- Renton, A.I., Dao, T.T., Johnstone, T. et al. Neurodesk: an accessible, flexible and portable data analysis environment for reproducible neuroimaging. Nat Methods (2024).

